Monitoring Gene Expression Changes As A Measure Of Cell Viability In 3D Cell Culture
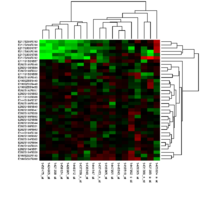
3-dimensional cell cultures are fast becoming the new standard for in vitro cell culture research owing to their efficient ability to mimic in vivo biology. To encourage this movement from 2D into 3D culture, proper quantitative methods need to be established to monitor the quality of cultures and measure relevant biological variables (1).
Gene expression profiles are already a well-established quantitative method of evaluating cell behaviour. The profiles are able to detect cellular gene expression changes upon external stimulation, and in response to environmental cues (2,3). The most common gene expression profiles targeted at measuring cell viability include measuring specific proteins and RNA sequences. However most of these methods involve the destruction of culture as micro arrays, RT-PCR or RNA sequencing methods requires lysis of cells. Additionally methods such as flow cytometry involve disintegration of culture. Immunohistochemical methods require fixing cultures, and other high throughput or transcriptome-wide technologies demands similar conditions. The primary reason behind such demands is the need to access RNA and proteins that are scarce in cells. Though they are indeed effective methods in monitoring cell viability in 2D cultures, their adoption in to 3D culture still remains a question (4).
Fluorescent probes
Several methods are now in development that would potentially allow quantitative evaluation of gene expression changes as a marker of cell viability, within in 3D culture. For example fluorescent probes, can enter the cells, and bind to targeted specific RNA sequences and give strong signals, without destruction of the culture. Though some limitations may occur in maintaining specificity, accuracy and uptake efficiency, these probes remain a viable option to use in long-term 3D cultures (4,5).
Gene Editing technologies
Novel alternative strategies include incorporating gene-editing technologies to couple a fluorescent reporter to the expression level of a gene of interest. It is a method with higher accuracy and precision. However, genetic modification adds an additional step, and can incur unwanted interactions or behavioural changes that may affect the overall culture quality (6-8). However, both fluorescent probes and gene editing technologies are well suited for monitoring gene expression changes over prolonged period of times, while preserving the distribution of cells within the culture. Furthermore they allow for evaluating the effect of different microenvironments on culture behaviour (1).
Among the potential avenues to increase the output of these assays, and make them more feasible to be used in 3D culturing, depend on using a combination of different readouts. This method was used for the visualization of complex cellular structures within 3D plant tissues (9); and with proper vital fluorescence-colorimetric methods it could be adapted to study gene expression in 3D cells. Despite its efficacy and precision, nearly all these methods mentioned above require substantial advances to be made in transcriptome/proteome wide analyses before being a viable option as a measure of cell viability in 3D cell cultures.
References
1. Cortesi M, Giordano E. Non-destructive monitoring of 3D cell cultures: new technologies and applications. PeerJ. 2022 May 12;10:e13338. doi: 10.7717/peerj.13338.
2. Picone G, Cappadone C, Pasini A, Lovecchio J, Cortesi M, Farruggia G, Lombardo M, Gianoncelli A, Mancini L, Ralf H M. 2020. Analysis of intracellular magnesium and mineral depositions during
osteogenic commitment of 3d cultured saos2 cells. International Journal of Molecular Sciences 21(7):2368.
3. Brady L, Gil da Costa RM, Coleman IM, Matson CK, Risk MC, Coleman RT, Nelson PS. 2020. A comparison of prostate cancer cell transcriptomes in 2D monoculture vs 3D xenografts identify consistent gene expression alterations associated with tumor microenvironments. The Prostate 80(6):491–499
4. Waylen LN, Nim HT, Martelotto LG, Ramialison M. 2020. From whole-mount to single-cell spatial assessment of gene expression in 3D. Communications Biology 3(1):1–11
5. Okamoto A. 2019. Next-generation fluorescent nucleic acids probes for microscopic analysis of intracellular nucleic acids. Applied Microscopy 49(1):1–7
6. He X, Chen H, Xu C, Fan J, Xu W, Li Y, Deng H, Shen J. 2020. Ratiometric and colorimetric fluorescent probe for hypochlorite monitor and application for bioimaging in living cells, bacteria and zebrafish. Journal of Hazardous Materials 388:122029
7. Di Blasi R, Zouein A, Ellis T, Ceroni F. 2021. Genetic toolkits to design and build mammalian synthetic systems. Trends in Biotechnology 39:P1004–P1018.
8. Koch B, Nijmeijer B, Kueblbeck M, Cai Y, Walther N, Ellenberg J. 2018. Generation and validation of homozygous fluorescent knock-in cells using CRISPR–Cas9 genome editing. Nature Protocols 13(6):1465
9. Ursache R, Andersen TG, Marhavy P, Geldner N. 2018. ` A protocol for combining fluorescent proteins with histological stains for diverse cell wall components. The Plant Journal 93(2):399–412 10. Wilkinson L, Friendly M. “The History of the Cluster Heat Map.
